8.3
Calculating heritability for threshold traits (diseases)
The calculations are based on disease frequencies in the general population (Population) and
among relatives (Indiv. related ) to a diseased animals. The relatives can be
(both parents diseased), first degree (father, mother or full sibs) or second
degree (half sibs or grand parents).Remember only one type of relatives can be analyzed at the same time.
Observe: The chi-square test calculation is based on: that the related ones is subtracted from the population numbers!
Example:
Enter your observed numbers and degree of relationship
and then press the Calculate button. The values in the (Degree of relationship) field should 1, .5 and .25 for the
three above mentioned cases.
You can also enter the frequencies but then the numbers shall be zeros
in the observed number fields.
Illustration of the case where both parents are affected are given below.
The program is initiated with the actual data for frequencies and the "Degree of relationship" is set to 1 (both parents diseased); to see
Press the Calculate button and see the results corresponding to the figure below.
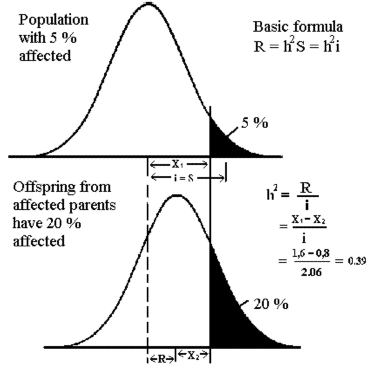
Remember in case of first or
second degree relative analyses with half or full sib, one affected individual per family is an
index case and should be subtracted from the group 'individual related'. So if there are 10
families with at least one affected in the population, 10 affected should be subtracted.
Questions:
In the table below is given observed numbers for disease frequencies in the
population and in first-degree relatives to diseased animals.
Normal Diseased Frequency
------------------------------------------
Population 980 20 0.020
First-degree
relative 140 10 0.066
------------------------------------------
Estimate the heritability.
What is the relative risk to be related to a diseased animal ?
Is the heritability statistically significant different from 0 with (p< 0.05) ?
Back to theory,
Back to theory (in Danish)
or Back to the other programs

